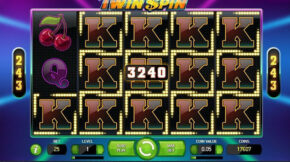
Nos últimos anos, os jogos de apostas online cresceram exponencialmente no Brasil, e um dos que mais tem chamado atenção é o Aviator. Simples, dinâmico e com potencial para altos lucros, esse jogo vem conquistando tanto apostadores iniciantes quanto os mais experientes. Mas será que é realmente possível ganhar com frequência no Aviator? Neste artigo, […]
-
 A IA Pode Ajudar Você a Ganhar no Cassino Online?maio 21, 2025
A IA Pode Ajudar Você a Ganhar no Cassino Online?maio 21, 2025

Jogar slots online é uma das formas mais populares de entretenimento em cassinos virtuais. Mas, para muitos jogadores, o sucesso ou a diversão de uma sessão não depende apenas da sorte, mas também do ambiente, do humor e do estado mental no qual se está ao jogar. Afinal, jogar em um momento adequado pode aumentar […]


Mais recente

Nos últimos anos, os jogos de apostas online cresceram exponencialmente no Brasil, e um dos que mais tem chamado atenção é o Aviator. Simples, dinâmico e com potencial para altos lucros, esse jogo vem conquistando tanto apostadores iniciantes quanto os mais experientes. Mas será que é realmente possível ganhar com frequência no Aviator? Neste artigo, […]

Entrar no mundo das apostas online pode ser emocionante, lucrativo e divertido — mas também pode ser arriscado se você não souber como se proteger. Seja em cassinos virtuais, sites de apostas esportivas ou slots online, é essencial começar com o pé direito e, principalmente, jogar com segurança. Neste guia, você vai descobrir tudo o […]

Nos últimos anos, as criptomoedas deixaram de ser um conceito obscuro de tecnologia para se tornarem um meio de pagamento real em diversos setores — e os cassinos online estão entre os que mais têm abraçado essa revolução. Bitcoin, Ethereum, Litecoin e outras moedas digitais estão mudando a forma como jogadores depositam, sacam e apostam […]

Jogar slots online é uma das formas mais populares de entretenimento em cassinos virtuais. Mas, para muitos jogadores, o sucesso ou a diversão de uma sessão não depende apenas da sorte, mas também do ambiente, do humor e do estado mental no qual se está ao jogar. Afinal, jogar em um momento adequado pode aumentar […]

A cultura asiática sempre foi rica em símbolos, histórias e mitos que fascinam o mundo inteiro. Nos últimos anos, muitos desses elementos migraram para o universo dos jogos online, especialmente nos slots de cassino e jogos de sorte. Um exemplo disso é a popular Série Fortune Ranqueada, que apresenta cinco animais icônicos da astrologia e […]